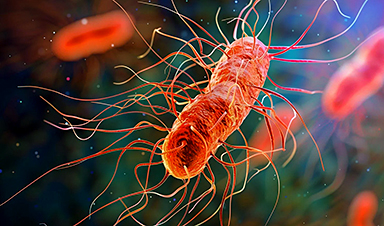
A mechanism used by bacteria to defend themselves could lead to the development of new antibiotics.
Princeton Engineering researchers have found a compound that can kill bacteria that cause incurable infections, with the potential to address the current drug-resistance crisis.
The compound, called cloacaenodin (chloa-say-nodin), is a short, slip-knotted chain of amino acids known as a lasso peptide, encoded by gut-dwelling bacteria as a defense mechanism. Peptides do all kinds of things in the body and have been used in a wide range of medical treatments. This peptide works by attacking rival bacteria, and it's a very potent killer, according to A. James Link, professor of chemical and biological engineering. If harnessed by science, it could be redirected to fight infections that are not treatable by today's medicines.
When released, the peptide hooks into a target cell's RNA-producing enzymes and shuts down basic cell functions. It targets an especially fearsome group of pathogens belonging to the genus Enterobacter, which the Center for Disease Control and Prevention (CDC) has identified as a primary driver in an accelerating global crisis: bacterial infections that increasingly do not respond to conventional antibiotics.
Link's research group has discovered several peptides in this same class — structured with a ring knotted to a tail that threads back down through the ring, like a lasso in a rodeo trick — that show promising antibacterial properties. He said cloacaenodin is unique because it can kill clinically relevant drug-resistant strains, making it a promising subject for antibiotic development. The finding also suggests his peptide-mining and synthetic biology techniques could reveal more antimicrobial compounds with strong drug-development potential, an essential step in quelling the growing superbug crisis.
"If it's made by one Enterobacter species, it's likely going to kill other species of Enterobacter. So it's this sort of guilt-by-association approach," Link said. This gives researchers a way to prioritize peptide-mining hits since peptides that are encoded in strains related to pathogens are more likely to have interesting bioactivity, he said.
An urgent need for new approaches
Ever since Anne Miller's fever broke on March 14, 1942, making her the first person ever saved by an antibiotic, humans have been simultaneously staving off deadly bacteria in the short run and saving millions of lives but also making infections harder to treat in the long run. Call it the law of unintended consequences. Some microbes have evolved rapidly to overwhelm our best efforts to destroy them.
The CDC has identified some Enterobacter species as a particularly urgent threat. Although harmless in the human gut, where they are common, when these bacteria enter the airways or urinary tract, they can cause serious infections. Many evade all known medicines, including a highly effective class of antibiotics known as carbapenems. So-called multi-drug resistance has ballooned over the past two decades. Untreatable infections now claim around a million lives each year, with that number projected to surpass cancer's death toll and reach 10 million per year by 2050, according to a 2019 United Nations report.
Market forces exacerbate the problem, according to the World Health Organization (WHO). Big pharmaceutical companies have strong financial incentives to pursue treatments for chronic conditions, where patient demand stretches over years. Because infections are treated in short finite intervals, profits from new antibiotics are relatively constrained. Adding to that, to slow drug-resistance dynamics, doctors tend to use newer drugs only after older drugs fail, leading to sluggish demand for small firms. And many new antibiotics don't present a clear advantage over cheaper, more familiar drugs. Over the past decade, several high-profile biotech startup companies with FDA-approved antibiotic treatments have collapsed under these economic conditions.
All of this has slowed the antibiotic-development pipeline to a trickle. The WHO has called the outlook "bleak." A recent report said that the "lack of diverse compounds suitable for bacterial treatment" and the "absence of new, suitable chemical matter to serve as leads for drug discovery is a major bottleneck in antibiotic discovery."
The non-profit organization CARB-X, run out of Boston University, has said developing new classes of antibiotics is the best strategy in addressing this urgent need. "You need a diversity of products," said CARB-X research and development chief Dr. Erin Duffy. "You need antibiotics — things that kill bacteria once you have an infection — and you need different classes, multiple classes." More than 20 classes of antibiotics were marketed in the two decades after Anne Miller's miraculous recovery. But since 1962 only two new antibiotic classes have made it to market, and neither treats the most resistant kinds of infections.
"It's one thing to kill bacteria," said Drew Carson, a fourth-year Ph.D. student in chemical and biological engineering and the paper's first author. "It's another thing to kill bacteria that can actually make people really sick."
A guilt-by-association approach
While cloacaenodin shows strong antibacterial properties, it's only the first of many steps to a new treatment. Determining a compound's safety is difficult and expensive, and moving from initial testing through the regulatory process takes a minimum of 10 years. Duffy said that, historically, some peptides have proven toxic to the kidneys, curbing their use in drugs. But peptides with bacterial-selective activity that don't harm animal cells will likely lack this toxicity, according to Link.
But this new compound shows promising antibacterial properties and the researchers have only just begun to consider what comes next. They plan to start by testing it in animal infection models to confirm that it can clear the infection and that it is safe for animal cells. More broadly, however, this compound's discovery suggests that Link and his team have developed a peptide-mining toolkit that will turn up many other interesting compounds in the future, and there is no telling where that will lead.
"The way that we find these peptides is by looking at the genome sequence of an organism," Link said. "If you give us any DNA sequence, we can very rapidly and very accurately figure out if there's a lasso peptide encoded within it. We also know about certain sequences within lasso peptides which means there's a good chance that they're antimicrobial. And that's how we homed in on this one."
Link said there are thousands of Enterobacter genome sequences that have been entered into scientific databases, and the lasso peptide his team discovered is found in only a handful. One of those organisms came from a hospital patient who had a lung infection. And because of his guilt-by-association approach to finding the peptide, they knew it would likely kill many related organisms that don't have the exact same genes.
"We tested it against a dozen or so strains and saw activity," Link said, referring to antibacterial activity. "But it potentially has activity against several hundred and maybe even thousands of these sequenced isolates of Enterobacter."
News
Ebola, hantavirus: Is the world prepared for the next pandemic?
Funding cuts to health research and a growing antivaccine movement are making it harder than ever to respond to viruses. The World Health Organization (WHO) has declared that an Ebola outbreak in Uganda and [...]
May 2026 Healthcare News and Trends: Market Signals That Matter
Artificial intelligence is dominating headlines, telehealth has settled into a new normal, and digital health continues to promise transformation. However, much of what is being discussed in healthcare today reflects potential rather than reality. [...]
Scientists Rewire Donor Stem Cells To Outsmart Aggressive Blood Cancers
Researchers have tested a gene-edited stem cell transplant designed to shield healthy blood-forming cells from powerful cancer-targeting immunotherapies. For patients with highly aggressive blood cancers, stem cell transplantation can offer a rare chance at [...]
Recent Digital Health Trends, Insights and News – May 2026
Last month marked continued progress as digital health moves into its next phase — from AI expanding into drug discovery and core infrastructure to new federal pathways accelerating device access and home-based care. Together, [...]
Cancer Mystery Solved: Scientists Discover How Melanoma Becomes “Immortal”
Scientists have uncovered a previously overlooked mechanism that may help melanoma cells become effectively “immortal.” Cancer cells face a major problem before they can become deadly: They have to figure out how to stop [...]
How Visual Neurons Organize Thousands of Synaptic Inputs
Summary: A new study uncovered the organizational rules that determine how neurons in the primary visual cortex process information. By imaging both the cell bodies (soma) and the individual synapses (on dendritic spines) of [...]
Scientists Just Found a Surprising Way To Destroy “Forever Chemicals”
Scientists have uncovered a new mechanism that may help break down highly persistent PFAS pollutants. PFAS have earned the nickname “forever chemicals” for a reason. These industrial compounds are so chemically durable that they [...]
Scientists Discover Cheap Material That Kills Deadly Superbugs
A new sulfur-rich antimicrobial polymer shows strong effectiveness against fungal and bacterial pathogens and may offer an affordable solution to antimicrobial resistance. Antimicrobial resistance is creating growing challenges for both healthcare and food production, [...]
What to Know About Cicada, or BA.3.2, the Latest SARS-CoV-2 Variant Under Monitoring
Like periodical cicadas, the insects for which it is nicknamed, SARS-CoV-2 Omicron subvariant BA.3.2 is only just beginning to emerge after lying low for an extended period since it first appeared. Although it was [...]
Scientists Say This Simple Supplement May Actually Reverse Heart Disease
Scientists in Japan say a common supplement may actually help “unclog” certain diseased heart arteries from the inside out. A simple food supplement sold in Japan may have helped reverse a dangerous form of [...]
New breakthrough against radiation: Korean Scientists create revolutionary shield with nanotechnology
Korean Scientists develop new nanotechnology material capable of reducing radiation impacts in space missions, hospitals, and power plants. The search for more efficient protection technologies in extreme environments has just gained an important advance. Korean [...]
Scientists Just Discovered the Hidden Trick That Keeps Your Cells Alive
A strange bead-like motion inside cells may be the secret to keeping their DNA—and health—in balance. Mitochondria are often described as the power plants of the cell because they produce the energy cells need [...]
Scientists Discover Stem Cells That Could Regrow Teeth and Bone
Scientists just uncovered the cellular “blueprint” that could one day let us regrow real teeth. Researchers at Science Tokyo have uncovered two distinct stem cell lineages that play a central role in forming tooth [...]
Scientists Uncover Fatal Weakness in “Zombie Cells” Linked to Cancer
A newly identified weakness in “zombie” cells may open the door to more precise cancer treatments by turning their own survival strategy against them. A new class of drugs takes advantage of a recently [...]
Bowel and Ovarian Cancers Are Dramatically Rising in Young Adults, Scientists Aren’t Sure Why
Cancer incidence is increasing, especially among younger adults, and current risk factors don’t fully account for the trend. Scientists suggest other underlying causes may be contributing. Cancer patterns in England are shifting in a [...]
New Immune Pathway Could Supercharge mRNA Cancer Vaccines
A surprising backup system in the immune response to mRNA vaccines may hold the key to more effective cancer treatments. The arrival of mRNA vaccines against SARS-CoV-2 in 2020 marked a turning point in the COVID-19 pandemic. Today, [...]