Molecular modeling suggests structural consequences of an early protein mutation that promoted viral transmission.
RIKEN researchers discovered that an early mutation (D614G) in the SARS-CoV-2 virus may have contributed to its rapid spread by altering the spike protein’s shape, improving the virus’s ability to adapt to human hosts. The finding could help inform the development of next-generation vaccines and antiviral drugs.
The rapid spread of COVID-19 may have been partly due to changes in the structure of the SARS-CoV-2 virus wrought by an early mutation in its genome, a detailed analysis by RIKEN researchers suggests. The finding could help inform the development of next-generation vaccines and antiviral drugs.
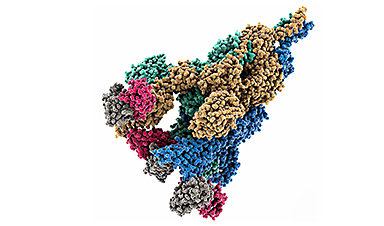
Yuji Sugita of the RIKEN Center for Computational Science (R-CCS) and Hisham Dokainish, who was at R-CCS at the time of the study, investigated the effect of mutations on viral structure. They did this by simulating the atomic positions of molecules found in different forms of the virus’s important spike protein—a tool coronaviruses use to bind and enter human cells.
They found that the substitution of a single amino acid altered this protein’s shape, helping SARS-CoV-2 to adapt to human hosts. This finding demonstrates how even tiny mutations—swapping a single amino acid in this case—can greatly affect protein dynamics.
To understand why the mutation proved so advantageous to the virus, the pair ran detailed simulations of the protein’s structure and stability. Their analysis—done using the RIKEN Fugaku supercomputer, one of the fastest in the world—revealed how the mutation (known as D614G) breaks an ionic bond with a second subunit of the Spike protein. It also changes the shape of a nearby loop structure, which alters the orientation of the entire protein, locking it into a form that makes it easier for the virus to enter cells (Fig. 1).
“A single and local change in an interaction within the molecule caused by a single mutation could affect the global structure of the spike protein,” explains Sugita, who is additionally affiliated with the RIKEN Center for Biosystems Dynamics Research. The resulting mutant proved better at replicating and transmitting between human hosts, and the D614Glineage quickly outcompeted its ancestral lineages and spread across the globe. It remains a fixture of every dominant variant that has followed.
Sugita’s team is now performing similar investigations of adaptive viral mutations that arose later in the course of the pandemic, including those found in the Omicron variant.
“Information obtained from our molecular dynamics simulations should help increase the opportunities for us to find effective drugs and other medicines,” he says.
News
New Once-a-Week Shot Promises Life-Changing Relief for Parkinson’s Patients
A once-a-week shot from Australian scientists could spare people with Parkinson’s the grind of taking pills several times a day. The tiny, biodegradable gel sits under the skin and releases steady doses of two [...]
Weekly injectable drug offers hope for Parkinson’s patients
A new weekly injectable drug could transform the lives of more than eight million people living with Parkinson's disease, potentially replacing the need for multiple daily tablets. Scientists from the University of South Australia [...]
Most Plastic in the Ocean Is Invisible—And Deadly
Nanoplastics—particles smaller than a human hair—can pass through cell walls and enter the food web. New research suggest 27 million metric tons of nanoplastics are spread across just the top layer of the North [...]
Repurposed drugs could calm the immune system’s response to nanomedicine
An international study led by researchers at the University of Colorado Anschutz Medical Campus has identified a promising strategy to enhance the safety of nanomedicines, advanced therapies often used in cancer and vaccine treatments, [...]
Nano-Enhanced Hydrogel Strategies for Cartilage Repair
A recent article in Engineering describes the development of a protein-based nanocomposite hydrogel designed to deliver two therapeutic agents—dexamethasone (Dex) and kartogenin (KGN)—to support cartilage repair. The hydrogel is engineered to modulate immune responses and promote [...]
New Cancer Drug Blocks Tumors Without Debilitating Side Effects
A new drug targets RAS-PI3Kα pathways without harmful side effects. It was developed using high-performance computing and AI. A new cancer drug candidate, developed through a collaboration between Lawrence Livermore National Laboratory (LLNL), BridgeBio Oncology [...]
Scientists Are Pretty Close to Replicating the First Thing That Ever Lived
For 400 million years, a leading hypothesis claims, Earth was an “RNA World,” meaning that life must’ve first replicated from RNA before the arrival of proteins and DNA. Unfortunately, scientists have failed to find [...]
Why ‘Peniaphobia’ Is Exploding Among Young People (And Why We Should Be Concerned)
An insidious illness is taking hold among a growing proportion of young people. Little known to the general public, peniaphobia—the fear of becoming poor—is gaining ground among teens and young adults. Discover the causes [...]
Team finds flawed data in recent study relevant to coronavirus antiviral development
The COVID pandemic illustrated how urgently we need antiviral medications capable of treating coronavirus infections. To aid this effort, researchers quickly homed in on part of SARS-CoV-2's molecular structure known as the NiRAN domain—an [...]
Drug-Coated Neural Implants Reduce Immune Rejection
Summary: A new study shows that coating neural prosthetic implants with the anti-inflammatory drug dexamethasone helps reduce the body’s immune response and scar tissue formation. This strategy enhances the long-term performance and stability of electrodes [...]
Scientists discover cancer-fighting bacteria that ‘soak up’ forever chemicals in the body
A family of healthy bacteria may help 'soak up' toxic forever chemicals in the body, warding off their cancerous effects. Forever chemicals, also known as PFAS (per- and polyfluoroalkyl substances), are toxic chemicals that [...]
Johns Hopkins Researchers Uncover a New Way To Kill Cancer Cells
A new study reveals that blocking ribosomal RNA production rewires cancer cell behavior and could help treat genetically unstable tumors. Researchers at the Johns Hopkins Kimmel Cancer Center and the Department of Radiation Oncology and Molecular [...]
AI matches doctors in mapping lung tumors for radiation therapy
In radiation therapy, precision can save lives. Oncologists must carefully map the size and location of a tumor before delivering high-dose radiation to destroy cancer cells while sparing healthy tissue. But this process, called [...]
Scientists Finally “See” Key Protein That Controls Inflammation
Researchers used advanced microscopy to uncover important protein structures. For the first time, two important protein structures in the human body are being visualized, thanks in part to cutting-edge technology at the University of [...]
AI tool detects 9 types of dementia from a single brain scan
Mayo Clinic researchers have developed a new artificial intelligence (AI) tool that helps clinicians identify brain activity patterns linked to nine types of dementia, including Alzheimer's disease, using a single, widely available scan—a transformative [...]
Is plastic packaging putting more than just food on your plate?
New research reveals that common food packaging and utensils can shed microscopic plastics into our food, prompting urgent calls for stricter testing and updated regulations to protect public health. Beyond microplastics: The analysis intentionally [...]