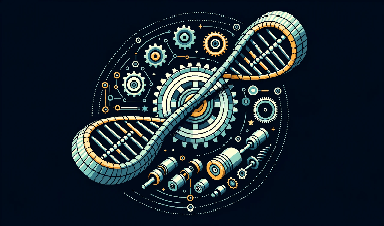
An international team of scientists has recently developed a novel type of nano engine made of DNA. It is driven by a clever mechanism and can perform pulsing movements. The researchers are now planning to fit it with a coupling and install it as a drive in complex nano machines. Their results have been published in the journal Nature Nanotechnology.
Šulc has used his group’s computer modeling tools to gain insights into design and operation of this leaf-spring nano engine. The structure is comprised of almost 14,000 nucleotides, which form the basic structural units of DNA.
“Being able to simulate motion in such a large nanostructure would be impossible without oxDNA, the computer model that our group uses for design and design of DNA nanostructures,” explains Šulc. “It is the first time that a chemically powered DNA nanotechnology motor has been successfully engineered. We are very excited that our research methods could help with studying it, and are looking forward to building even more complex nanodevices in the future.”
This novel type of engine is similar to a hand grip strength trainer that strengthens your grip when used regularly. However, the motor is around one million times smaller. Two handles are connected by a spring in a V-shaped structure.
In a hand grip strength trainer, you squeeze the handles together against the resistance of the spring. Once you release your grip, the spring pushes the handles back to their original position. “Our motor uses a very similar principle,” says professor Michael Famulok from the Life and Medical Sciences (LIMES) Institute at the University of Bonn. “But the handles are not pressed together but rather pulled together.”
The researchers have repurposed a mechanism without which there would be no plants or animals on Earth. Every cell is equipped with a sort of library. It contains the blueprints for all types of proteins that each cell needs to perform its function. If the cell wants to produce a certain type of protein, it orders a copy from the respective blueprint. This transcript is produced by the enzymes called RNA polymerases.
RNA polymerases drive the pulsing movements
The original blueprint consists of long strands of DNA. The RNA polymerases move along these strands and copy the stored information letter by letter.
“We took an RNA polymerase and attached it to one of the handles in our nanomachine,” explains Famulok, who is also a member of the transdisciplinary research areas “Life & Health” and “Matter” at the University of Bonn.
“In close proximity, we also strained a DNA strand between the two handles. The polymerase grabs on to this strand to copy it. It pulls itself along the strand and the non-transcribed section becomes increasingly smaller. This pulls the second handle bit by bit towards the first one, compressing the spring at the same time.”
The DNA strand between the handles contains a particular sequence of letters shortly before its end. This so-called termination sequence signals to the polymerase that it should let go of the DNA. The spring can now relax again and moves the handles apart. This brings the start sequence of the strand close to the polymerase and the molecular copier can start a new transcription process: The cycle then repeats.
“In this way, our nanomotor performs a pulsing action,” explains Mathias Centola from the research group headed by professor Famulok, who carried out a large proportion of the experiments.
An alphabet soup serves as fuel
This motor also needs energy just like any other type of motor. It is provided by the “alphabet soup” from which the polymerase produces the transcripts. Every one of these letters (in technical terminology: nucleotides) has a small tail consisting of three phosphate groups—a triphosphate.
In order to attach a new letter to an existing sentence, the polymerase has to remove two of these phosphate groups. This releases energy which it can use for linking the letters together. “Our motor thus uses nucleotide triphosphates as fuel,” says Famulok. “It can only continue to run when a sufficient number of them are available.”
The researchers were able to demonstrate that the motor can be easily combined with other structures. This should make it possible for it to, for example, wander across a surface—similar to an inchworm that pulls itself along a branch in its own characteristic style.
“We are also planning to produce a type of clutch that will allow us to only utilize the power of the motor at certain times and otherwise leave it to idle,” explains Famulok. In the long term, the motor could become the heart of a complex nanomachine. “However, there is still a lot of work to be done before we reach this stage.”
Šulc’s lab is highly interdisciplinary and applies broadly the methods of statistical physics and computational modeling to problems in chemistry, biology and nanotechnology. The group develops new multiscale models to study interactions between biomolecules, particularly in the context of design and simulations of DNA and RNA nanostructures and devices.
“Just as complex machines in our everyday use—planes, cars and chips in electronics—require sophisticated computer-aided design tools to make sure they perform a desired function, there is a pressing need to have access to such methods in the molecular sciences.”
Professor Tijana Rajh, director of the School of Molecular Sciences, said, “Petr Šulc and his group are doing extremely innovative molecular science, using the methods of computational chemistry and physics to study DNA and RNA molecules in the context of biology as well as nanotechnology. Our younger faculty members in the School of Molecular Sciences have an extraordinary record of achievement, and Professor Šulc is an exemplar in this regard.”
Bio-nanotechnology
DNA and RNA are the basic molecules of life. They fulfill many functions, including information storage and information transfer in living cells. They also have promising applications in the field of nanotechnology where designed DNA and RNA strands are used to assemble nanoscale structures and devices.
As Šulc explains, “It is a little bit like playing with Lego blocks except that each Lego block is only a few nanometers (a millionth of a millimeter) in size, and instead of putting each block into the place where it should go, you put them inside a box and shake it randomly until only the desired structure comes out.”
This process is called self-assembly, and Šulc and his colleagues use computational modeling and design software to come up with the building blocks that reliably assemble into the shape one wants at nanoscale resolution.
“The promising applications of this field include diagnostics, therapeutics, molecular robotics, and building of new materials,” says Šulc.
“My lab has developed the software to design these blocks, and we work closely with experimental groups at ASU as well as other universities in the U.S. and Europe. It is exciting seeing our methods used to design and characterize nanostructures of increasing complexity, as the field progresses and we achieve new advanced designs and successfully operate them at nanoscale.”
More information: A rhythmically pulsing leaf-spring DNA-origami nanoengine that drives a passive follower, Nature Nanotechnology (2023). DOI: 10.1038/s41565-023-01516-x. www.nature.com/articles/s41565-023-01516-x
Journal information: Nature Nanotechnology
News
Cancer Mystery Solved: Scientists Discover How Melanoma Becomes “Immortal”
Scientists have uncovered a previously overlooked mechanism that may help melanoma cells become effectively “immortal.” Cancer cells face a major problem before they can become deadly: They have to figure out how to stop [...]
How Visual Neurons Organize Thousands of Synaptic Inputs
Summary: A new study uncovered the organizational rules that determine how neurons in the primary visual cortex process information. By imaging both the cell bodies (soma) and the individual synapses (on dendritic spines) of [...]
Scientists Just Found a Surprising Way To Destroy “Forever Chemicals”
Scientists have uncovered a new mechanism that may help break down highly persistent PFAS pollutants. PFAS have earned the nickname “forever chemicals” for a reason. These industrial compounds are so chemically durable that they [...]
Scientists Discover Cheap Material That Kills Deadly Superbugs
A new sulfur-rich antimicrobial polymer shows strong effectiveness against fungal and bacterial pathogens and may offer an affordable solution to antimicrobial resistance. Antimicrobial resistance is creating growing challenges for both healthcare and food production, [...]
What to Know About Cicada, or BA.3.2, the Latest SARS-CoV-2 Variant Under Monitoring
Like periodical cicadas, the insects for which it is nicknamed, SARS-CoV-2 Omicron subvariant BA.3.2 is only just beginning to emerge after lying low for an extended period since it first appeared. Although it was [...]
Scientists Say This Simple Supplement May Actually Reverse Heart Disease
Scientists in Japan say a common supplement may actually help “unclog” certain diseased heart arteries from the inside out. A simple food supplement sold in Japan may have helped reverse a dangerous form of [...]
New breakthrough against radiation: Korean Scientists create revolutionary shield with nanotechnology
Korean Scientists develop new nanotechnology material capable of reducing radiation impacts in space missions, hospitals, and power plants. The search for more efficient protection technologies in extreme environments has just gained an important advance. Korean [...]
Scientists Just Discovered the Hidden Trick That Keeps Your Cells Alive
A strange bead-like motion inside cells may be the secret to keeping their DNA—and health—in balance. Mitochondria are often described as the power plants of the cell because they produce the energy cells need [...]
Scientists Discover Stem Cells That Could Regrow Teeth and Bone
Scientists just uncovered the cellular “blueprint” that could one day let us regrow real teeth. Researchers at Science Tokyo have uncovered two distinct stem cell lineages that play a central role in forming tooth [...]
Scientists Uncover Fatal Weakness in “Zombie Cells” Linked to Cancer
A newly identified weakness in “zombie” cells may open the door to more precise cancer treatments by turning their own survival strategy against them. A new class of drugs takes advantage of a recently [...]
Bowel and Ovarian Cancers Are Dramatically Rising in Young Adults, Scientists Aren’t Sure Why
Cancer incidence is increasing, especially among younger adults, and current risk factors don’t fully account for the trend. Scientists suggest other underlying causes may be contributing. Cancer patterns in England are shifting in a [...]
New Immune Pathway Could Supercharge mRNA Cancer Vaccines
A surprising backup system in the immune response to mRNA vaccines may hold the key to more effective cancer treatments. The arrival of mRNA vaccines against SARS-CoV-2 in 2020 marked a turning point in the COVID-19 pandemic. Today, [...]
Scientists Discover “Molecular Switch” That Fuels Alzheimer’s Brain Inflammation
A newly identified trigger of brain inflammation could offer a fresh target for slowing Alzheimer’s progression. The brain has its own built-in immune system that identifies threats and responds to them. In Alzheimer’s disease, growing evidence [...]
Molecular Manufacturing: The Future of Nanomedicine – New book from NanoappsMedical Inc.
This book explores the revolutionary potential of atomically precise manufacturing technologies to transform global healthcare, as well as practically every other sector across society. This forward-thinking volume examines how envisaged Factory@Home systems might enable the cost-effective [...]
Forgotten Medicinal Plant Shows Promise in Fighting Dangerous Superbugs
A traditional medicinal plant, tormentil, shows promise against antibiotic-resistant bacteria in laboratory tests. Its compounds work by limiting bacterial growth and boosting antibiotic performance. Before the development of modern antibiotics, plant-based remedies were commonly [...]
NanoMedical Brain/Cloud Interface – Explorations and Implications. A new book from Frank Boehm
New book from Frank Boehm, NanoappsMedical Inc Founder: This book explores the future hypothetical possibility that the cerebral cortex of the human brain might be seamlessly, safely, and securely connected with the Cloud via [...]