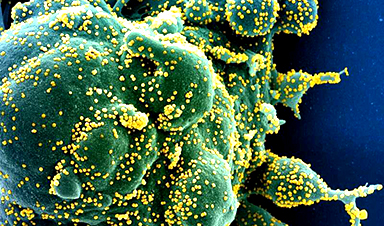
The COVID-19 pandemic has prompted considerable investigation into how the SARS-CoV-2 Spike protein attaches to a human cell during the infection process, as this knowledge is useful in designing vaccines and therapeutics. Now, a team of scientists has discovered additional locations on the Spike protein that may not only help to explain how certain mutations make emerging variants more infectious but also could be used as additional targets for therapeutic intervention.
“Significant research is underway to examine how the receptor binding domain (RBD) at the tip of the club-shaped SARS-CoV-2 Spike protein attaches to an ACE2 receptor on a human cell, but little is known about the other changes that occur in the Spike protein as a result of this attachment,” said Ganesh Anand, associate professor of chemistry, Penn State. “We have uncovered ‘hotspots’ further down on the Spike protein that are critical for SARS-CoV-2 infection and may be novel targets beyond the RBD for therapeutic intervention.”
Anand and his colleagues used a process, called amide hydrogen-deuterium exchange mass spectrometry (HDXMS), to visualize what happens when the SARS-CoV-2 Spike protein binds to an ACE2 receptor. HDXMS uses heavy water or deuterium oxide (D2O), a naturally occurring, non-radioactive isotope of water formed from heavy hydrogen or deuterium, as a probe for mapping proteins. In this case, the team placed SARS-CoV-2 Spike protein and ACE2 receptors in heavy water and obtained footprints of ACE2 on the Spike protein.
“If you put the Spike protein and ACE2 receptor into a solution that’s made with D2O, the surfaces and more floppy regions on both proteins will more readily exchange hydrogens for deuterium, compared to their interiors,” said Anand. “And footprints of each protein on the binding partner can be readily identified from areas where you see little deuterium and only detect normal hydrogen.”
Using this technique, the team determined that binding of the Spike protein and ACE2 receptor is necessary for furin-like proteases—a family of human enzymes—that act to snip off the tip, called the S1 subunit, of the Spike protein, which is the next step in the virus’s infection of the cell. The findings published on Feb. 8 in the journal eLife.
Image Credit: NIH/NIAID
Post by Amanda Scott, NA CEO. Follow her on twitter @tantriclens
Thanks to Heinz V. Hoenen. Follow him on twitter: @HeinzVHoenen
News
Why More People in Their 30s Are Suddenly Getting Colon Cancer
A major Swiss study found that colorectal cancer is becoming increasingly common in adults under 50, even as rates decline in older age groups. Researchers in Switzerland have identified a concerning trend: while colorectal [...]
Researchers Compare MS Models to Human Tissue in Search for Better Therapies
Researchers identified key differences between two widely used multiple sclerosis models, showing how each can better study myelin damage, immune responses, and repair. The findings may improve efforts to develop treatments that restore lost [...]
Scientists Discover Genetic “Off Switch” That Supercharges CAR T Cells Against Cancer
A new study reveals a possible way to make CAR T-cell therapy more durable and effective by targeting a single gene-regulating protein. CAR T-cell therapy is widely seen as a breakthrough in personalized cancer [...]
New Vitamin B12-Based Therapy Could Change How Brain Cancer Is Treated
Researchers have identified a vitamin B12–based compound that appears capable of crossing the blood–brain barrier and selectively accumulating in glioblastoma tissue. For decades, one of the biggest problems in brain cancer treatment has had [...]
Simple Fiber Supplement Cuts Knee Arthritis Pain in Just 6 Weeks, Study Finds
A daily inulin supplement may help reduce knee osteoarthritis pain while revealing a possible link between gut health, muscle function, and pain sensitivity. For millions of people living with knee osteoarthritis, managing chronic pain [...]
This Common Vitamin May Help Stop Prediabetes From Turning Into Diabetes
Vitamin D may help prevent type 2 diabetes in people with specific genetic variations, offering a possible path toward personalized diabetes prevention. More than 40% of U.S. adults have prediabetes, a condition in which [...]
Ebola, hantavirus: Is the world prepared for the next pandemic?
Funding cuts to health research and a growing antivaccine movement are making it harder than ever to respond to viruses. The World Health Organization (WHO) has declared that an Ebola outbreak in Uganda and [...]
May 2026 Healthcare News and Trends: Market Signals That Matter
Artificial intelligence is dominating headlines, telehealth has settled into a new normal, and digital health continues to promise transformation. However, much of what is being discussed in healthcare today reflects potential rather than reality. [...]
Scientists Rewire Donor Stem Cells To Outsmart Aggressive Blood Cancers
Researchers have tested a gene-edited stem cell transplant designed to shield healthy blood-forming cells from powerful cancer-targeting immunotherapies. For patients with highly aggressive blood cancers, stem cell transplantation can offer a rare chance at [...]
Recent Digital Health Trends, Insights and News – May 2026
Last month marked continued progress as digital health moves into its next phase — from AI expanding into drug discovery and core infrastructure to new federal pathways accelerating device access and home-based care. Together, [...]
Cancer Mystery Solved: Scientists Discover How Melanoma Becomes “Immortal”
Scientists have uncovered a previously overlooked mechanism that may help melanoma cells become effectively “immortal.” Cancer cells face a major problem before they can become deadly: They have to figure out how to stop [...]
How Visual Neurons Organize Thousands of Synaptic Inputs
Summary: A new study uncovered the organizational rules that determine how neurons in the primary visual cortex process information. By imaging both the cell bodies (soma) and the individual synapses (on dendritic spines) of [...]
Scientists Just Found a Surprising Way To Destroy “Forever Chemicals”
Scientists have uncovered a new mechanism that may help break down highly persistent PFAS pollutants. PFAS have earned the nickname “forever chemicals” for a reason. These industrial compounds are so chemically durable that they [...]
Scientists Discover Cheap Material That Kills Deadly Superbugs
A new sulfur-rich antimicrobial polymer shows strong effectiveness against fungal and bacterial pathogens and may offer an affordable solution to antimicrobial resistance. Antimicrobial resistance is creating growing challenges for both healthcare and food production, [...]
What to Know About Cicada, or BA.3.2, the Latest SARS-CoV-2 Variant Under Monitoring
Like periodical cicadas, the insects for which it is nicknamed, SARS-CoV-2 Omicron subvariant BA.3.2 is only just beginning to emerge after lying low for an extended period since it first appeared. Although it was [...]
Scientists Say This Simple Supplement May Actually Reverse Heart Disease
Scientists in Japan say a common supplement may actually help “unclog” certain diseased heart arteries from the inside out. A simple food supplement sold in Japan may have helped reverse a dangerous form of [...]