t typically takes many years of experiments to develop a new medicine. Although vaccines to protect against disease from the novel coronavirus are starting to reach clinics around the world, patients and doctors will still need treatments to manage COVID-19 symptoms for some time.
At Pacific Northwest National Laboratory (PNNL), computational biologists, structural biologists, and analytical chemists are using their expertise to safely accelerate the design step of the COVID-19 drug discovery process.
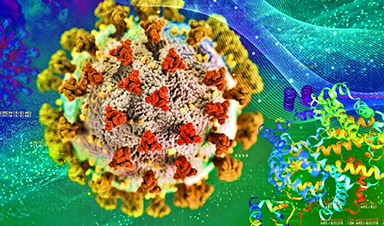
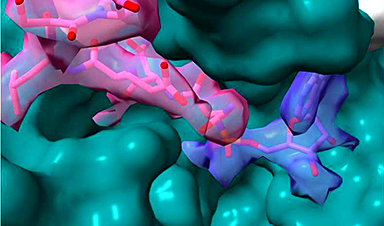
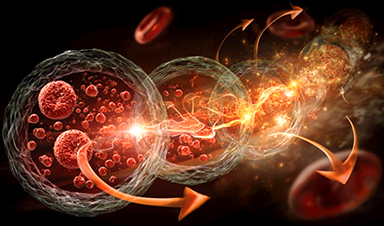
Rather than finding a new drug by trial and error, scientists are taking the three-dimensional structures of proteins from the novel coronavirus and using computer modeling and machine learning to identify a unique molecule that best fits inside a binding pocket on a protein’s surface. Ideally, that molecule clogs the viral protein and prevents it from functioning.
“Drug research and development is a complex, costly, and time-consuming process, particularly considering the majority of molecules advanced from the design phase fail in clinical trials,” said PNNL computational data scientist Neeraj Kumar. “Computer-based screening incorporates chemical information during the design process to increase a drug candidate’s potential for success in clinical testing.”
Developing an approach to speed drug discovery during this pandemic could also reveal new design steps that might be useful during the next outbreak.
Clogging coronavirus proteins
There are almost 30 different proteins in this novel coronavirus that are potential targets for COVID-19 drug discovery. Combine that with millions of molecules that are potential drug candidates, and the possibilities for matching molecules to specific proteins are mind-boggling.
To narrow the options towards molecules with potential to become medicines, Kumar and his team first use molecular docking to virtually screen libraries of known molecules and regulatory-approved drugs. Ones that fit in the binding pocket of a particular coronavirus protein make the short list for the next step of the process: testing the fit with actual proteins and molecules.
Experimental scientists then combine the molecules on this short list with purified coronavirus protein and “weigh them” with native mass spectrometry to see if the protein picked up the molecule. This technique measures interactions between the protein and the molecules and can confirm the predicted binding.
Image Credit: Timothy Holland | Pacific Northwest National Laboratory
Post by Amanda Scott, NA CEO. Follow her on twitter @tantriclens
Thanks to Heinz V. Hoenen. Follow him on twitter: @HeinzVHoenen
News
Repurposed drugs could calm the immune system’s response to nanomedicine
An international study led by researchers at the University of Colorado Anschutz Medical Campus has identified a promising strategy to enhance the safety of nanomedicines, advanced therapies often used in cancer and vaccine treatments, [...]
Nano-Enhanced Hydrogel Strategies for Cartilage Repair
A recent article in Engineering describes the development of a protein-based nanocomposite hydrogel designed to deliver two therapeutic agents—dexamethasone (Dex) and kartogenin (KGN)—to support cartilage repair. The hydrogel is engineered to modulate immune responses and promote [...]
New Cancer Drug Blocks Tumors Without Debilitating Side Effects
A new drug targets RAS-PI3Kα pathways without harmful side effects. It was developed using high-performance computing and AI. A new cancer drug candidate, developed through a collaboration between Lawrence Livermore National Laboratory (LLNL), BridgeBio Oncology [...]
Scientists Are Pretty Close to Replicating the First Thing That Ever Lived
For 400 million years, a leading hypothesis claims, Earth was an “RNA World,” meaning that life must’ve first replicated from RNA before the arrival of proteins and DNA. Unfortunately, scientists have failed to find [...]
Why ‘Peniaphobia’ Is Exploding Among Young People (And Why We Should Be Concerned)
An insidious illness is taking hold among a growing proportion of young people. Little known to the general public, peniaphobia—the fear of becoming poor—is gaining ground among teens and young adults. Discover the causes [...]
Team finds flawed data in recent study relevant to coronavirus antiviral development
The COVID pandemic illustrated how urgently we need antiviral medications capable of treating coronavirus infections. To aid this effort, researchers quickly homed in on part of SARS-CoV-2's molecular structure known as the NiRAN domain—an [...]
Drug-Coated Neural Implants Reduce Immune Rejection
Summary: A new study shows that coating neural prosthetic implants with the anti-inflammatory drug dexamethasone helps reduce the body’s immune response and scar tissue formation. This strategy enhances the long-term performance and stability of electrodes [...]
Scientists discover cancer-fighting bacteria that ‘soak up’ forever chemicals in the body
A family of healthy bacteria may help 'soak up' toxic forever chemicals in the body, warding off their cancerous effects. Forever chemicals, also known as PFAS (per- and polyfluoroalkyl substances), are toxic chemicals that [...]
Johns Hopkins Researchers Uncover a New Way To Kill Cancer Cells
A new study reveals that blocking ribosomal RNA production rewires cancer cell behavior and could help treat genetically unstable tumors. Researchers at the Johns Hopkins Kimmel Cancer Center and the Department of Radiation Oncology and Molecular [...]
AI matches doctors in mapping lung tumors for radiation therapy
In radiation therapy, precision can save lives. Oncologists must carefully map the size and location of a tumor before delivering high-dose radiation to destroy cancer cells while sparing healthy tissue. But this process, called [...]
Scientists Finally “See” Key Protein That Controls Inflammation
Researchers used advanced microscopy to uncover important protein structures. For the first time, two important protein structures in the human body are being visualized, thanks in part to cutting-edge technology at the University of [...]
AI tool detects 9 types of dementia from a single brain scan
Mayo Clinic researchers have developed a new artificial intelligence (AI) tool that helps clinicians identify brain activity patterns linked to nine types of dementia, including Alzheimer's disease, using a single, widely available scan—a transformative [...]
Is plastic packaging putting more than just food on your plate?
New research reveals that common food packaging and utensils can shed microscopic plastics into our food, prompting urgent calls for stricter testing and updated regulations to protect public health. Beyond microplastics: The analysis intentionally [...]
Aging Spreads Through the Bloodstream
Summary: New research reveals that aging isn’t just a local cellular process—it can spread throughout the body via the bloodstream. A redox-sensitive protein called ReHMGB1, secreted by senescent cells, was found to trigger aging features [...]
AI and nanomedicine find rare biomarkers for prostrate cancer and atherosclerosis
Imagine a stadium packed with 75,000 fans, all wearing green and white jerseys—except one person in a solid green shirt. Finding that person would be tough. That's how hard it is for scientists to [...]
Are Pesticides Breeding the Next Pandemic? Experts Warn of Fungal Superbugs
Fungicides used in agriculture have been linked to an increase in resistance to antifungal drugs in both humans and animals. Fungal infections are on the rise, and two UC Davis infectious disease experts, Dr. George Thompson [...]