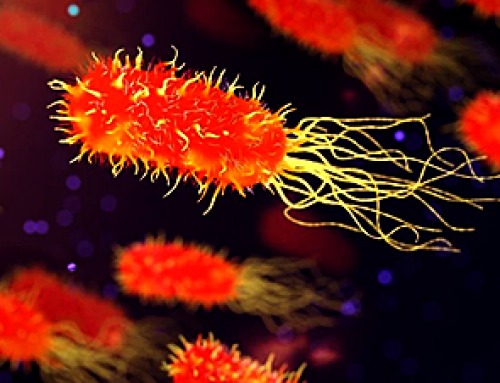
A decades-old mystery of how natural antimicrobial predatory bacteria are able to recognize and kill other bacteria may have been solved, according to new research.
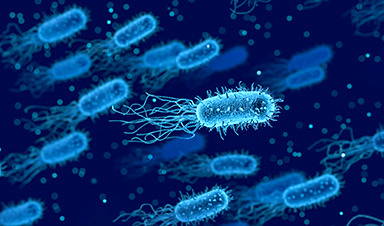
In a study published in Nature Microbiology, researchers from the University of Birmingham and the University of Nottingham have discovered how natural antimicrobial predatory bacteria, called Bdellovibrio bacterivorous, produce fiber-like proteins on their surface to ensnare prey.
This discovery may enable scientists to use these predators to target and kill problematic bacteria that cause issues in healthcare, food spoilage, and the environment.
Professor of Structural Biology at the University of Birmingham, Andrew Lovering said: "Since the 1960s Bdellovibrio bacterivorous has been known to hunt and kill other bacteria by entering the target cells and eating them from the inside before later bursting out. The question that had stumped scientists was 'how do these cells make a firm attachment when we know how varied their bacterial targets are?'"
Professor Lovering and Professor Liz Sockett, from the School of Life Sciences at the University of Nottingham, have been collaborating in this field for almost 15 years. The breakthrough came when Sam Greenwood an undergraduate student, and Asmaa Al-Bayati, a PhD student in the Sockett lab, discovered that the Bdellovibrio predators lay down a sturdy vesicle (a "pinched-off" part of the predator cell envelope) when invading their prey.
The Role of Vesicles and Fibers in Predation
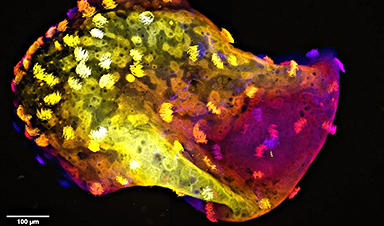
Professor Liz Sockett explained: "The vesicle creates a kind of airlock or keyhole allowing Bdellovibrio entry into the prey cell. We were then able to isolate this vesicle from the dead prey, which is a first in this field. The vesicle was analyzed to reveal the tools used during the preceding event of predator/prey contact. We thought of it as a bit like a locksmith leaving the pick, or key, as evidence, in the keyhole.
"By looking at the vesicle contents, we discovered that because Bdellovibrio doesn't know which bacteria it will meet, it deploys a range of similar prey recognition molecules on its surface, creating lots of different 'keys' to 'unlock' lots of different types of prey."
Advanced Molecular Analysis and Engineering
The researchers then undertook an individual analysis of the molecules, demonstrating that they form long fibers, approximately ten times longer than common globular proteins. This allows them to operate at a distance and "feel" for prey in the vicinity.
In total, the labs counted 21 different fibers. Researchers Dr Simon Caulton, Dr Carey Lambert, and Dr Jess Tyson worked on how they operated both at the cellular and molecular levels. They were supported by fibre gene-engineering by Paul Radford and Rob Till. The team then began to attempt linking a particular fiber to a particular prey-surface molecule. Finding out which fiber matches which prey, could enable an engineering approach that sees bespoke predators targeting different types of bacteria.
Professor Lovering continued: "Because the predator strain we were looking at comes from the soil it has a wide killing range, making this identification of these fiber and prey pairs very difficult. However, on the fifth attempt to find the partners we discovered a chemical signature on the outside of prey bacteria that was a tight fit to the fibre tip. This is the first time a feature of Bdellovibrio has been matched to prey selection."
Scientists in this field will now be able to use these discoveries to ask which fiber set is used by the different predators they study and potentially attribute these to specific prey. Improving understanding of these predator bacteria could enable their usage as antibiotics, to kill bacteria that degrade food, or ones that are harmful to the environment.
Professor Lovering concluded: "We know that these bacteria can be helpful, and by fully understanding how they operate and find their prey, it opens up a world of new discoveries and possibilities."
Reference: "Bdellovibrio bacteriovorus uses chimeric fibre proteins to recognize and invade a broad range of bacterial hosts" by Simon G. Caulton, Carey Lambert, Jess Tyson, Paul Radford, Asmaa Al-Bayati, Samuel Greenwood, Emma J. Banks, Callum Clark, Rob Till, Elisabete Pires, R. Elizabeth Sockett and Andrew L. Lovering, 4 January 2024, Nature Microbiology.
DOI: 10.1038/s41564-023-01552-2
The research was funded by the Wellcome Trust Investigator in Science Award (209437/Z/17/Z).
News
Scientists Discover “Molecular Switch” That Fuels Alzheimer’s Brain Inflammation
A newly identified trigger of brain inflammation could offer a fresh target for slowing Alzheimer’s progression. The brain has its own built-in immune system that identifies threats and responds to them. In Alzheimer’s disease, growing evidence [...]
Molecular Manufacturing: The Future of Nanomedicine – New book from NanoappsMedical Inc.
This book explores the revolutionary potential of atomically precise manufacturing technologies to transform global healthcare, as well as practically every other sector across society. This forward-thinking volume examines how envisaged Factory@Home systems might enable the cost-effective [...]
Forgotten Medicinal Plant Shows Promise in Fighting Dangerous Superbugs
A traditional medicinal plant, tormentil, shows promise against antibiotic-resistant bacteria in laboratory tests. Its compounds work by limiting bacterial growth and boosting antibiotic performance. Before the development of modern antibiotics, plant-based remedies were commonly [...]
NanoMedical Brain/Cloud Interface – Explorations and Implications. A new book from Frank Boehm
New book from Frank Boehm, NanoappsMedical Inc Founder: This book explores the future hypothetical possibility that the cerebral cortex of the human brain might be seamlessly, safely, and securely connected with the Cloud via [...]
New Research Finds Shocking Link Between Chili Peppers and Cancer
If you love spicy food, you are not alone. But scientists are taking a closer look at whether eating a lot of chili peppers could affect your cancer risk. Could your love of spicy [...]
New book from Nanoappsmedical Inc. – Global Health Care Equivalency
A new book by Frank Boehm, NanoappsMedical Inc. Founder. This groundbreaking volume explores the vision of a Global Health Care Equivalency (GHCE) system powered by artificial intelligence and quantum computing technologies, operating on secure [...]
Scientists Create “Neurobots” – Living Machines With Their Own Nervous Systems
Neurobots—xenobots with neurons—show self-organized nervous systems and enhanced behaviors, revealing new insights into how biology builds functional structures. In 2020, researchers at Tufts University developed tiny living structures known as xenobots using frog cells. These microscopic organisms [...]
Our books now available worldwide!
Online Sellers other than Amazon, Routledge, and IOPP Indigo Global Health Care Equivalency in the Age of Nanotechnology, Nanomedicine and Artifcial Intelligence Global Health Care Equivalency In The Age Of Nanotechnology, Nanomedicine And Artificial [...]
Amazonian Chocolate Could Become the Next Superfood, Scientists Say
New research into Amazonian cocoa reveals that its value may extend beyond flavor alone. Chocolate from the Amazon is already known worldwide for its distinctive taste, but new research suggests it may offer even [...]
Nanobody repairs misfolded CFTR inside cells, boosting function in cystic fibrosis
A tiny antibody component could fundamentally transform the treatment of cystic fibrosis: For the first time, researchers have succeeded in developing a so-called nanobody that penetrates directly into human cells and can repair the [...]
20-Year Study Finds Daily Multivitamins Don’t Extend Lifespan
A large, decades-long study of over 390,000 U.S. adults challenges a widespread assumption about daily multivitamins. Multivitamins are a daily habit for millions of Americans, often taken with the expectation that they will extend [...]
Novel Investment Paradigms for Regenerative Healthcare Ecosystems
Introduction The transition toward regenerative healthcare ecosystems—anchored in wellness optimization, disease prevention, eradication strategies, and healthy longevity—necessitates a structural reconfiguration of capital architectures, governance models, and incentive design. Regenerative healthcare, by definition, transcends episodic [...]
What If Consciousness Exists Beyond Your Brain
Scientists still don’t know how consciousness emerges from the brain. New ideas suggest it may not emerge at all, but instead be a basic feature of reality. Is consciousness produced by the brain, or [...]
Scientists Discover Way To Treat Lung Cancer and Its Deadly Side Effect Together
A new approach using lipid nanoparticles to deliver genetic material is showing promise in tackling two major challenges in lung cancer at once.Researchers at Oregon State University have designed a new way to tackle two of [...]
Saunas Activate Your Immune System
A brief sauna session may quietly mobilize the immune system. A sauna session may do more than raise your heart rate and body temperature. A new study from Finland found that it also briefly [...]
Why music from your youth still has such an intense effect years later: A psychological perspective
You're driving, and suddenly a familiar song fills the air. Before you even know it, a wave of emotions comes over you – not just memories, but a deep, almost physical feeling. This powerful [...]